INDIANAPOLIS—Researchers at Indiana University School of Medicine have discovered alternative gene splicing, which occurs during gene expression, can impact a person’s risk of alcohol use disorder (AUD). They recently published their findings in Molecular Psychiatry.
“AUD is a common and complex genetic disorder that happens people experience problems related to excessive alcohol consumption,” said Rudong Li, PhD, a postdoctoral fellow in the YunLong Liu, PhD Laboratory and lead author of the paper. “This discovery has revealed a novel perspective about AUD and opens up new possibilities for finding new therapeutics.”
Alternative splicing of RNA controls the flow of genetic information from DNA to gene expression and is known to be associated with many complex diseases, especially neurological or brain disorders. The team took a statistical genetics approach to identify exons on the genes that are skipped and could contribute to AUD risk. Using models in the laboratory, they found 27 exon skipping events that affect AUD risk.
“This is the first time that we’ve seen how the exon inclusion on specific genes can potentially lead to addiction,” said Yunlong Liu, PhD, director of the Center for Computational Biology and Bioinformatics and senior author of the study. “We used novel computational methods for understanding the roles of alternative splicing in complex disease by innovatively combining transcriptomics data from post-mortem brain tissues with genome-wide association studies (GWAS) data on disease traits.”
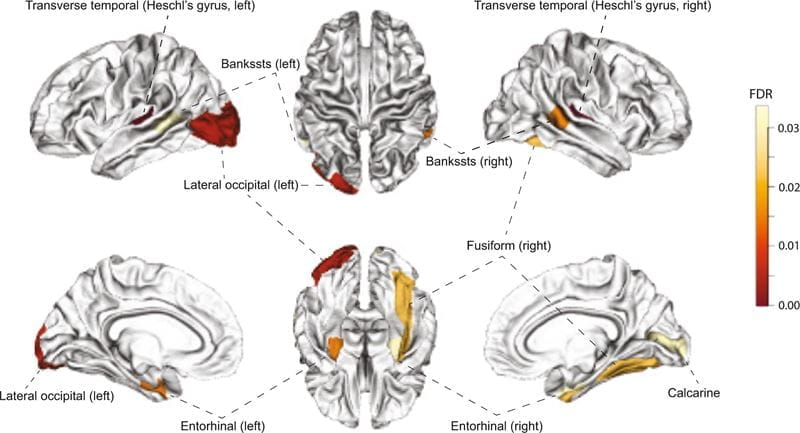
Li says future research could target the novel genes, or regions of special interest of the genes, to further understand molecular mechanisms in complex diseases, including AUD and other substance use disorders, and potentially develop new therapeutics.
“This discovery could change people’s understanding of AUD and the science behind it,” said Li.
In addition to Li and Liu, other study authors include Jill Reiter, PhD; Andy Chen, MS, PhD; Steven Chen; Tatiana Foroud, PhD; Howard Edenberg, PhD; and Dongbing Lai, MS, PhD.
Read the full publication in Molecular Psychiatry.
Learn more about alcohol research at IU School of Medicine.
About IU School of Medicine
IU School of Medicine is the largest medical school in the United States and is annually ranked among the top medical schools in the nation by U.S. News & World Report. The school offers high-quality medical education, access to leading medical research and rich campus life in nine Indiana cities, including rural and urban locations consistently recognized for livability.